Abstract
Background
SARS-CoV-2 has been detected not only in respiratory secretions, but also in stool collections. Here were sought to identify SARS-CoV-2 by enrichment next-generation sequencing (NGS) from fecal samples, and to utilize whole genome analysis to characterize SARS-CoV-2 mutational variations in COVID-19 patients.
Results
Study participants underwent testing for SARS-CoV-2 from fecal samples by whole genome enrichment NGS (n = 14), and RT-PCR nasopharyngeal swab analysis (n = 12). The concordance of SARS-CoV-2 detection by enrichment NGS from stools with RT-PCR nasopharyngeal analysis was 100%. Unique variants were identified in four patients, with a total of 33 different mutations among those in which SARS-CoV-2 was detected by whole genome enrichment NGS.
Conclusion
These results highlight the potential viability of SARS-CoV-2 in feces, its ongoing mutational accumulation, and its possible role in fecal–oral transmission. This study also elucidates the advantages of SARS-CoV-2 enrichment NGS, which may be a key methodology to document complete viral eradication.
Trial registration ClinicalTrials.gov, NCT04359836, Registered 24 April 2020, https://clinicaltrials.gov/ct2/show/NCT04359836?term=NCT04359836&draw=2&rank=1).
Background
December 2019, the virus, which would cause a worldwide pandemic, was first identified in the city of Wuhan, China [1]. In January 2020, it was implicated in various pneumonia cases, and was rapidly isolated from a bronchoalveolar lavage sample, analyzed via next-generation sequencing (NGS), and identified to be a novel betacoronavirus [2, 3], the same family of viruses responsible for severe acute respiratory syndrome (SARS) and middle east respiratory syndrome (MERS) [4,5,6]. This new virus, named the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) on February 11, 2020 by the International Committee on the Taxonomy of Viruses (ICTV), has now been implicated in over 13.4 million cases worldwide, and over 580,000 deaths [7, 8]. Over 138,000 of these deaths have been in the United States alone.
Early diagnosis of SARS-CoV-2 has been challenging and may contribute to its high mortality rates. Due to the similarities between SARS-CoV-2 and SARS-CoV, a similar diagnostic approach utilizing real time polymerase chain reaction (RT-PCR) from nasopharyngeal swabs has been adopted as the standard of care [9]. Contributing to the spread of the disease, has been the viral load being below the threshold of detection in some individuals [10]. Additionally, acquired mutations may assist in its evasion of detection from specifically targeted PCR primers [11]. Further, samples collected soon after infection, or after symptoms have resolved, have resulted in high false negative rates. Samples collected in these timeframes have been shown to have false-negative rates as high as 33% [12]. The exact rate of false positives has not been established for SARS-CoV-2, however in a study by Cohen and Kessel, the rate was extrapolated based on known false-positive rates from other viral RT-PCR tests. The mean false-positive rate in the study was found the be 3.2%, ranging from 0% to 16.7%, and an interquartile range of 0.8 to 4.0% [13].
Symptoms of SARS-CoV-2 infection cover a broad range including fever, malaise, body aches, chills, headaches, cough, and shortness of breath, as well as gastrointestinal symptoms such as diarrhea, nausea, and vomiting [14, 15]. Research from SARS and MERS has shown that coronaviruses can be present in stool of infected patients, and that the fecal–oral route may be a mode of transmission. Wang et al. in 2005 demonstrated that SARS-CoV was not only present in stool samples collected from patients, but also in the wastewater of two hospitals [16]. In review of this data, with inclusion of SARS-CoV-2, a meta-analysis by Parasa et al. found that 40% of patients with positive RT-PCR for SARS-CoV-2 from nasopharyngeal swabs or respiratory secretions were also found to be positive from stool samples [17]. This data was supported by two recent studies documenting positive SARS-CoV-2 fecal samples from “recovered” COVID-19 patients with negative nasopharyngeal swabs [18, 19].
In view of the large percentage of SARS-CoV-2 detectible by RT-PCR in stools of infected patients, we sought to identify the presence of the SARS-CoV-2 by NGS of fecal samples from symptomatic study participants positive for SARS-CoV-2 by nasopharyngeal sample RT-PCR, in addition to asymptomatic individuals (with or without prior nasopharyngeal sample RT-PCR). We also aimed to execute whole genome analysis to characterize SARS-CoV-2 mutational variations to identify potentially significant nucleotide changes.
Results
We evaluated the results from patients that had their stool samples tested by whole genome enrichment NGS, and their nasopharyngeal swabs tested by RT-PCR for the presence of SARS-CoV-2. Of the 14 study participants, ten were symptomatic and tested positive for SARS-CoV-2 by RT-PCR, two asymptomatic individuals tested negative, and two other asymptomatic individuals did not undergo RT-PCR testing (Table 1). While all patients were asked to collect stools at baseline, the samples were collected from 2 to 38 days after RT-PCR testing. Patients 5, 7, and 13, who all tested PCR positive by nasopharyngeal swab, were treated by their primary care physicians. Patients 5 and 7, were treated with Hydroxychloroquine (HCQ), Azithromycin (Zpack), vitamin C (3000 mg), vitamin D (3000 IU), and zinc (50 mg) for 10 days. Stools from patients 5 and 7 were collected 5 and 6 days respectively after therapy regimens were initiated, at which time, both patients reported symptom clearance. Similarly, after positive nasopharyngeal swab, patient 13 began treatment consisting of high dosages of vitamin C (up to 10,000 mg), vitamin D (up to 3000 IU), and zinc (up to 50 mg). This patient’s stool was collected 4 days after beginning the 10-day treatment regimen. Again, dosage changes were not recorded; however, symptom clearance was noted before fecal collection.
Table 1 Symptoms and SARS-CoV-2 testing results
The concordance of SARS-CoV-2 detection by enrichment NGS from stools among positive non-treated patients tested by RT-PCR nasopharyngeal analysis was 100% (7/7). Patient 8, who did not undergo nasopharyngeal analysis, tested positive for SARS-CoV-2 by NGS. The three patients (5, 7, 13) that received treatment and were asymptomatic prior to providing fecal samples, were tested negative by NGS. Asymptomatic patients 2 and 9, who tested negative by nasopharyngeal swab, were also negative by NGS, as was asymptomatic patient 14.
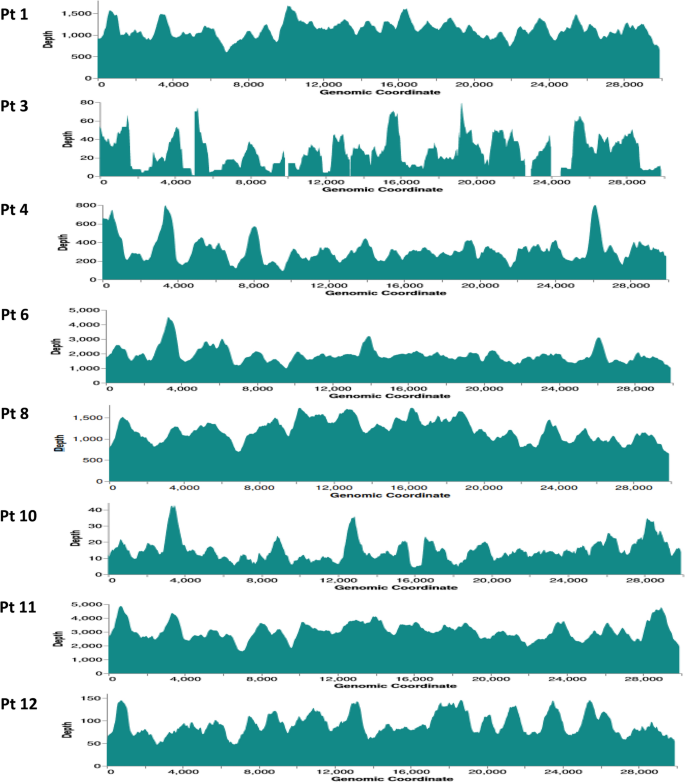
The enrichment NGS analysis of fecal samples collected from RT-PCR-positive patients demonstrated mean read depths of 1129.8x, 31.7x, 318.6x, 1924.6x, 1206.7x, 15.6x, 3075.3x, and 92.7x, against the SARS-CoV-2 genome in patients 1, 3, 4, 6, 8, 10, 11, and 12, respectively. The ATCC SARS-CoV-2 served as a positive control in this study, with 7856.3x mean read depth. The sequencing read mapping results are depicted in Fig. 1 (read depths are denoted on the y-axis and specific genomic coordinates on the x-axis). At 10x minimum read depth, all but two samples achieved a 100% breadth of coverage of the Wuhan-Hu-1 SARS-CoV-2 reference genome (Table 2). In patients 3 and 10, 35% and 59% of genome completeness were achieved, respectively.
Fig. 1
Mapping outcomes of patient NGS data against the SARS-CoV-2 Wuhan-Hu-1 (MN90847.3) complete genome. The x-axis depicts the genomic coordinates and the y-axis represents the read depth at specific loci
Table 2 Enrichment NGS metrics
The total number of SARS-CoV-2 mapped reads for patients 1, 3, 4, 6, 8, 10, 11, and 12 were 465,645, 5984, 131,582, 793,603, 496,852, 5929, 1,270,734, and 38,256 respectively (ATCC SARS-CoV-2 positive control, 94,693,754 reads).
Following alignment and mapping of SARS-CoV-2, patient genomes were compared to the Wuhan-Hu-1 (MN90847.3) SARS-CoV-2 reference genome via One Codex’s bioinformatics pipeline to identify mutational variations (see Methods for details). This analysis identified nucleotide variants at positions nt241 (C → T) and nt23403 (A → G) across all positive patients, and variants at positions nt3037 (C → T) and nt25563 (G → T) in seven of the eight patients (Table 3). Interestingly, patients 8, 11, and 12 harbored the same set of variants, as did patients 4 and 6 (who were kindreds). Unique variants not identified in any of the other individuals were detected in patients 1, 3, 6, and 10, with patient 3 harboring the most distinct SARS-CoV-2 genome with eight unique variants, followed by patient 1 with seven. Collectively, there were thirty-three different mutations among the patients in which SARS-CoV-2 was detected by whole genome enrichment NGS. One limitation to the variant analysis was that the One Codex SARS-CoV2 pipeline does not identify putative amino acids changes. As a result, Table 3 only captures nucleotide level resolution, making it difficult to assess mutations effect and evolutionary relationships.
Table 3 SARS-CoV-2 genomic positions, variant changes, and frequencies across the positive patient cohort
Discussion
Coronaviridae is a family of enveloped, single-stranded, positive-sense RNA viruses [20, 21]. The total length of the genome is 30 Kb, consisting of a 5′-terminal noncoding region, an open reading frame (ORF) 1a/b-coding region, an S region encoding the spike glycoprotein (S protein), an E region encoding the envelope protein (E protein), an M region encoding the membrane protein (M protein), an N region encoding the nucleocapsid protein (N protein), and a 3′-terminal noncoding region [22,23,24,25]. Among them, the poly protein encoded in the ORF1a/b region of the nonstructural protein can be cut by 3CLpro and PLpro of the virus to form RNA-dependent RNA polymerase and helicase, which guides the replication, transcription, and translation of the virus genome. The M and E proteins are involved in the formation of the envelope, while the N protein is involved in assembly. The spike protein binds to the receptor of the host cell and confers specificity for viral invasion into susceptible cells.
Once decoded, the SARS-CoV-2 genome was found to share high sequence identity with the bat coronavirus, BatCoV RaTG13 (96.2%) (2). Upon further investigation, it was discovered that SARS-CoV-2 harbored significant sequence homology with the viruses responsible for SARS and MERS, with a notable exception found in the receptor binding domain (RBD) [26]. Shang et al. elucidated the RBD structure of the human ACE2 receptor (angiotensin-converting enzyme 2) [27], demonstrating that the replacement of several residues within the protein caused it to have a much more compact hydrophobic pocket. This change increased the binding affinity of SARS-CoV-2 to ACE2 as compared to SARS-CoV. While this has contributed to its greater virulence, it also represents a potential therapeutic target [28]. This concept was detailed in the mini-review by Yang and Shen, wherein they proposed that SARS-CoV-2 may be susceptible to the inhibitory effect of chloroquine (CQ), a lysosomotropic agent, via its accumulation in the acidic organelles [29]. The therapeutic effect of CQ may be a result of its ability to neutralize the endosome-lysosomal acidic pH and block the protease activity necessary for viral entry [30]. This could possibly be evidenced by the HCQ treated patients in this study that appeared to have cleared the virus having tested negative by enrichment NGS for SARS-CoV-2 while having testing positive by nasopharyngeal analysis prior to treatment. While the small sample size and lack of randomization preclude any statistical significance for this finding, the sequencing test demonstrated the capacity to detect SARS-Cov-2 in stool, as well as to detect viral clearance, which warrants further study. We make no assertion whether the viral clearance was due to treatment received or was natural clearance.
Two recent studies that support targeting the endocytic pathway and autophagy as therapeutic strategies are by Arshad et al. and Lagier et al., both of which show that HCQ was associated with a significant reduction of in-hospital mortality compared to those not receiving HCQ [31, 32]. Numerous other studies have also reported the efficacy of CQ and/or HCQ in various treatment regimens for COVID-19 [33,34,35,36,37,38,39]. Adding to the complexity of COVID-19 treatment and prevention, is that SARS-CoV-2 appears to be mutating at an alarming rate, as reported in the Icelandic study which identified the presence of 291 sequence variants that were not present in the Global Initiative on Sharing All Influenza Data (GISAID) reference database as of March 22 (11).
Although previous studies have identified SARS-CoV-2 in fecal collections by RT-PCR [40, 41], this is the first to our knowledge, to report whole genome sequencing (WGS) of SARS-CoV-2 from stool samples. Herein we were able to identify SARS-CoV-2 in patients that tested positive by nasopharyngeal swab RT-PCR analysis and obtained complete viral genomes in 6 out of 8 NGS-positive patients. The overall homology among the genomes was high (99.97%), with variations identified in the ORF regions 1a, 1b, S, 3a, 8, and N. Of particular interest, was the adenine to guanine change in the S protein at position nt23403 which converts aspartic acid to glycine (D → G). Although the significance of this variation is unclear, it warrants further investigation to understand its effect on spike glycoprotein ACE2 binding and virulence. The conversions of glycine to arginine (nt28883) and proline to arginine (nt29364) in the nucleoprotein also necessitate further examination. While enrichment NGS is both costly and time consuming, these striking results highlight the potential viability of SARS-CoV-2 in feces, its possible role in transmission, and may accurately document complete eradication of the virus.
Conclusion
Next generation sequencing identified the SARS-CoV-2 whole genome sequence in 100% of patients with positive nasopharyngeal RT-PCR and did not detect it in asymptomatic post-treatment patients, or those with negative RT-PCR. Of notable interest, was that patient 1 still tested positive for SARS-CoV-2 by NGS from stool, 38 days after positive nasopharyngeal RT-PCR test. This information suggests that the virus may linger for longer than anticipated in the GI tract and warrants further longitudinal investigation to understand if the virus is viable and/or transmissible via fecal material, and if so, for how long is it contagious in this capacity. Collectively, these results highlight the importance of metagenomic analysis of the SARS-CoV-2 viral genome, and present an alternative diagnostic methodology that may help with viral identification, and tracking of its evolutionary progression through the population, as well as its clearance.
Methods
Study participants (n = 14) underwent testing for SARS-CoV-2 from fecal samples by whole genome enrichment NGS. Following fecal collection in Zymo Research Shield Fecal Collection Tubes, stool samples were transported to the laboratory where RNA was extracted utilizing the Qiagen Allprep Power Fecal Kit from 200 μl of stool material that was suspended in the DNA/RNA Shield stabilization solution present in the Zymo collection vials. Zymo’s DNA/RNA Shield is designed to preserve RNA genetic integrity and prevent degradation at ambient temperature for > 1 month, or > -20 °C indefinitely. All samples were adhered to manufacture’s specifications to assure sample stability. Included throughout sample processing was the SARS-CoV-2 positive control from ATCC (Heat-inactivated SARS-CoV-2, VR-1986HK; strain 2019-nCoV/USA-WA1/2020), and the no template control (NTC) to monitor extraneous nucleic acid contamination. Following purification, all available viral RNA was reverse transcribed (New England Biolabs NEBNext 1st and 2nd Strand Synthesis Modules), library prepped (Illumina Nextera Flex for Enrichment and IDT for Illumina-Nextera DNA UD Indexes Set), enriched (Ilumina Respiratory Virus Oligo Panel), and sequenced with Illumina’s NextSeq 500/550 High-Output v2.5 300 cycle kit on the Illumina NextSeq 550 System. Run set-up parameters on the NextSeq Control Software (Illumina Local Run Manager) included paired-end sequencing set to 76 cycles with both Index 1 and 2 at 10 bp (base pair). The 76 bp selection is appropriate due to the size of the baits in the Illumina Virus Oligo Panel. Although the sequencing kit utilized has the capacity to sequence at 2 × 150 bp, selection above 76 cycles would begin to sequence into the adapters, negatively impacting NGS quality metrics. Sequencing acceptance criteria were a Q-score (AQ30) ≥ 75%, cluster density between 120 and 240 K/mm2, and clusters passing filter (PF%) ≥ 80%. Following successful NGS QC, sequences were then mapped utilizing the minimap2 sequencing alignment tool in One Codex’s SARS-CoV-2 bioinformatics analysis pipeline. SARS-CoV-2 positive samples were further analyzed for mutational variants that differed from the reference genome. The complete analysis pipeline for SARS-CoV-2 is open source and available at http://github.com/onecodex/sars-co-v-2. A detailed description of the bioinformatics methods is available at http://docs.onecodex.com/en/articles/3793936-covid-10-sequencing-analysis. SARS-CoV-2 genome sequencing data (in patients from which complete viral genomes were obtained) are deposited in NBCI’s GenBank database (Accession IDs: MW425856, MW425855, MW425854, MW425853, MW425852, MW425851 for patients 1, 4, 6, 8, 11, and 12 respectively). The SARS-Cov-2 amino acid changes reported in the discussion section were manually analyzed utilizing the NCBI database (http://ncbi.nlm.nih.gov/gene). Of the 14 study participants, 12 also had their nasopharyngeal swabs tested for SARS-CoV-2 by RT-PCR.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
- Huang C, Wang Y, Li X, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan. China Lancet. 2020;395:497–506.
- Article CAS PubMed Google Scholar
- Zhou P, Yang X-L, Wang X-G, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–3.
- Article CAS PubMed PubMed Central Google Scholar
- Zhu N, Zhang D, Wang W, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020;382:727–33.
- Article CAS PubMed PubMed Central Google Scholar
- Drosten C, Günther S, Preiser W, et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med. 2003;348:1967–76.
- Article CAS PubMed Google Scholar
- Ksiazek TG, Erdman D, Goldsmith CS, et al. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med. 2003;348:1953–66.
- Article CAS PubMed Google Scholar
- Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus ADME, Fouchier RAM. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med. 2012;367:1814–20.
- Article CAS PubMed Google Scholar
- “Coronavirus.” World Health Organization, World Health Organization, 15 July 2020, www.who.int/emergencies/diseases/novel-coronavirus-2019.
- Johns Hopkins University. Coronavirus Resource Center. COVID-19 Dashboard by the Center for Systems Science and Engineering (CSSE) at Johns Hopkins University (JHU). 15 July 2020, www.coronavirus.jhu.edu/map.html.
- Wang D, Hu B, Hu C, et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus_infected Pneumonia in Wuhan. China JAMA. 2020. https://doi.org/10.1001/jama.2020.1585.
- Article PubMed Google Scholar
- Tahamtan A, Ardebili A. Real-Time RT-PCR in COVID-19 detection: issues affecting the results. Expert Rev Molecular Diagnostics. 2020;20(5):453–4. https://doi.org/10.1080/14737159.2020.1757437.
- Article CAS Google Scholar
- Gudbjartsson D, et al. Spread of SARS-CoV-2 in the Icelandic Population. N Engl J Med. 2020;382(24):2302–15. https://doi.org/10.1056/nejmoa2006100.
- Article CAS PubMed Google Scholar
- Wikramaratna P, et al. Estimating False-Negative Detection Rate of SARS-CoV-2 by RT-PCR. MedRxiv. 2020. https://doi.org/10.1101/2020.04.05.20053355.
- Article Google Scholar
- Cohen A, Kessel B. False positives in reverse transcription PCR testing for SARS-CoV-2. MedRxiv. 2020. https://doi.org/10.1101/2020.04.26.20080911.
- Article PubMed PubMed Central Google Scholar
- Report of the WHO-China joint mission on coronavirus disease 2019 (COVID-19); https://www.who.int/docs/default-source/coronaviruse/who-china-joint-mission-on-covid-19-final-report.pdf Disease. 2020 2020
- Kakodkar, Pramath, et al. A Comprehensive Literature Review on the Clinical Presentation, and Management of the Pandemic Coronavirus Disease 2019 (COVID-19). Cureus, 2020, doi: https://doi.org/10.7759/cureus.7560.
- Wang X-W. Excretion and detection of SARS coronavirus and its nucleic acid from digestive system. World J Gastroenterol. 2005;11(28):4390. https://doi.org/10.3748/wjg.v11.i28.4390.
- Article CAS PubMed PubMed Central Google Scholar
- Parasa S, et al. Prevalence of gastrointestinal symptoms and fecal viral shedding in patients with coronavirus disease 2019. JAMA Network Open. 2020. https://doi.org/10.1001/jamanetworkopen.2020.11335.
- Article PubMed PubMed Central Google Scholar
- Zhang B, Liu S, Dong Y, Zhang L, Zong Q, Zou Y, Zhang S. Positive rectal swabs in young patients recovered from coronavirus disease 2019 (COVID-19). J Infect. 2020;13:37. https://doi.org/10.1016/j.jinf.2020.04.023.
- Article CAS Google Scholar
- Chen L, Lou J, Bai Y, et al. COVID-19 disease with positive fecal and negative pharyngeal and sputum viral tests. Am J Gastroenterol. 2020. https://doi.org/10.14309/ajg.0000000000000610.
- Article PubMed PubMed Central Google Scholar
- Weiss SR, Navas-Martin S. Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol Mol Biole Rev. 2005;69:635–64.
- Article CAS Google Scholar
- Scoeman D, Fielding BC. Coronavirus enveloped protein: current knowledge. Virol J. 2019;16:69.
- Article Google Scholar
- Zumla A, Chan J, Azhar E, et al. Coronaviruses—drug discovery and therapeutic options. Nat Rev Drug Discov. 2016;15:327–47. https://doi.org/10.1038/nrd.2015.37.
- Article CAS PubMed PubMed Central Google Scholar
- Zhao L, et al. Antagonism of the interferon-induced OAS-RNase L pathway by murine coronavirus ns2 protein is required for virus replication and liver pathology. Cell Host Microbe. 2012;11(6):607–16. https://doi.org/10.1016/j.chom.2012.04.011.
- Article CAS PubMed PubMed Central Google Scholar
- Belshaw R, et al. The evolution of genome compression and genomic novelty in RNA viruses. Genome Res. 2007;17(10):1496–504. https://doi.org/10.1101/gr.6305707.
- Article CAS PubMed PubMed Central Google Scholar
- Fehr A, Perlman S. Coronaviruses: an overview of their replication and pathogenesis. Coronaviruses Methods Molecular Biol. 2015. https://doi.org/10.1007/978-1-4939-2438-7_1.
- Article Google Scholar
- Li F. Structure, Function, and Evolution of Coronavirus Spike Proteins. Annual Rev Virol. 2016;3(1):237–61. https://doi.org/10.1146/annurev-virology110615-042301.
- Article CAS Google Scholar
- Shang J, et al. Structural basis of receptor recognition by SARS-CoV-2. Nature. 2020;581(7807):221–4. https://doi.org/10.1038/s41586-020-2179-y.
- Article CAS PubMed PubMed Central Google Scholar
- Yang N, Shen H. Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. Int J Biol Sci. 2020;16(10):1724–31. https://doi.org/10.7150/ijbs.45498.
- Article CAS PubMed PubMed Central Google Scholar
- Degtyarev M, De Maziere A, Orr C, Lin J, Lee BB, Tien JY, et al. Akt inhibition promotes autophagy and sensitizes PTEN-null tumors to lysosomotropic agents. J Cell Biol. 2008;183:101–16.
- Article CAS PubMed PubMed Central Google Scholar
- Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell res. 2020.
- Arshad S, Kilgore P, Chaudhry Z, et al. Treatment with Hydroxychloroquine, Azithromycin, and Combination in Patients Hospitalized with COVID-19. Int J Infect Dis. 2020. https://doi.org/10.1016/j.jijid.2020.06.099.
- Article PubMed PubMed Central Google Scholar
- Lagier JC, Matthieu M, Gautret P, Colson P, Cortaredona S, et al. Outcomes of 3,737 COVID-19 patients treated with hydroxychloroquine/azithromycin and other regimens in Marseille, France: A retrospective analysis. Travel Med Infect Dis. 2020. https://doi.org/10.1016/j.tmaid.2020.101791.
- Article PubMed PubMed Central Google Scholar
- Gao J, Tian Z, Yang X. Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci Trends. 2020;14(1):72–3.
- Article CAS PubMed Google Scholar
- Gautret P, Lagier JC, Parola P, et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int J Antimicrob Agents 2020;105949.
- Million M, Lagier JC, Gautret P, et al. Early treatment of COVID-19 patients with hydroxychloroquine and azithromycin: A retrospective analysis of 1061 cases in Marseille, 29 France. Travel Med Infect Dis. 2020;5:101738. https://doi.org/10.1016/j.tmaid.2020.101738.
- Article Google Scholar
- Yu B, Wang DW, Li C. Hydroxychloroquine application is associated with a decreased mortality in critically ill patients with COVID-19. medRxiv. 2020. doi:https://doi.org/10.1101/2020.04.27.20073379
- Chen Z, Hu J, Zhang Z, Jiang S, Han S, Yan D, Zhuang R, Hu B, Zhang Z. Efficacy of hydroxychloroquine in patients with COVID-19: results of a randomized clinical trial. medRxiv. 2020. doi: https://doi.org/10.1101/2020.03.22.20040758.
- Huang M, Tang T, Pang P, Li M, Ma R, Lu J, Shu J, You Y, Chen B, Liang J, Hong Z, Chen H, Kong L, Qin D, Pei D, Xia J, Jiang S, Shan H. Treating COVID-19 with Chloroquine. 2020. J Mol Cell Biol. doi: https://doi.org/10.1093/jmcb/mjaa014.
- Geleris J, Sun Y, Platt J, et al. Observational Study of Hydroxychloroquine in Hospitalized Patients with Covid-19. N Engl J Med. 2020.
- Chen Y, et al. The Presence of SARS-CoV-2 RNA in the Feces of COVID-19 Patients. J Med Virol. 2020;92(7):833–40. https://doi.org/10.1002/jmv.25825.
- Article CAS PubMed Google Scholar
- Wu Y, et al. Prolonged Presence of SARS-CoV-2 Viral RNA in Fecal Samples. Lancet Gastroenterol Hepatol. 2020;5(5):434–5. https://doi.org/10.1016/s2468-1253(20)30083-2.
- Article PubMed PubMed Central Google Scholar
Acknowledgements
We thank Dr. Alan Miller at Alta Pharmaceuticals for contributing to providing patient samples. We thank Nick Greenfield and Christopher Smith at One Codex for providing bioinformatic support.
Funding
This study was funded by ProgenaBiome.
Author information
Authors and Affiliations
- ProgenaBiome, Ventura, CA, United States
- Andreas Papoutsis, Jordan Daniels, Skylar Steinberg, Brad Barrows & Sabine Hazan
- Centre for Digestive Diseases, Sydney, Australia
- Thomas Borody & Siba Dolai
- Ventura Clinical Trials, Ventura, CA, United States
- Sabine Hazan
Contributions
Conceptualization, AP and SH; Methodology, AP and SH; Investigation, AP and SH Writing-original draft, AP; Writing-review and editing, AP, TB, SD, JD, SS, BB, and SH Supervision, SH; Funding acquisition, SH. All authors read and approved the final manuscript.
Corresponding author
Correspondence to Andreas Papoutsis.
Ethics declarations
Ethics approval and consent to participate
This study was approved by the institutional review board of Advarra. Consent to participate in this study from each patient was agreed via signed informed consent.
Consent for publication
Agreed to by patients on the signed informed consent, demonstrating that the patient understood the procedures and the purpose of the study.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Papoutsis, A., Borody, T., Dolai, S. et al. Detection of SARS-CoV-2 from patient fecal samples by whole genome sequencing. Gut Pathog 13, 7 (2021). https://doi.org/10.1186/s13099-021-00398-5
- Received24 July 2020
- Accepted12 January 2021
- Published30 January 2021
- DOIhttps://doi.org/10.1186/s13099-021-00398-5
Share this article
Anyone you share the following link with will be able to read this content:
Provided by the Springer Nature SharedIt content-sharing initiative
Keywords
Gut Harmony: Unlocking Autism Relief Through Microbiome Balance”
“Microbial Medicine: How Gut Health Affects Autism Symptoms”
“Healing from Within: The Gut-Brain Connection in Autism”
“Navigating the Microbiome Maze: Autism and Gut Wellness”
“Microbes as Allies: Restoring Balance for Autism Relief”
“Autism Unraveled: Microbiome Insights and Symptom Alleviation”
“Gut Check: Balancing Microbiota for Autism Wellness”
“Microbiome Magic: Relieving Autism Symptoms Naturally”
“From Dysbiosis to Hope: Gut Health and Autism Management”
“Microbial Symphony: Improving Autism Well-Being via the Gut”
Read More: Mastering LinkedIn Marketing: LinkedIn marketing Edmonton